
Click on image for
Ninth Day Intro
 

"Initial Genetic
Characterization of
the 1918 'Spanish'
Influenza Virus,"
Science.
Volume 275,
pages 1793-1796
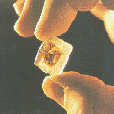
Wax slice containing
autopsied lung sample
from 1918 victim�
|
![[Interview conducted by Leonard Crane in May of 1998]](images/ninthday/Conducted.gif)
Introduction:
In March of 1997 the influenza research community was startled by the appearance
of a remarkable 3-page report� in the journal Science. In the paper
Dr. Jeffery Taubenberger and colleagues, of the Armed Forces Institute of Pathology,
announced that they had recovered and analyzed fragments of the RNA genome of the
virus responsible for the infamous 1918 Spanish Flu
pandemic.
The shattered genes of the virus were retrieved from lung-tissue samples fixed by
formalin, preserved in paraffin wax, and stored for the last 80 years at the National
Tissue Repository, located within the Walter Reed Army Medical Center, in Maryland.
The warehouse contains autopsy specimens dating back to the time of the Civil War.
Because of RNA's extreme delicacy, the team's successful extraction and sequencing
of the genetic material has both delighted and tantalized those seeking to understand
the mysteries of this historically and epidemiologically important influenza strain.
Full sequencing of the 1918 genome is expected to take years, although one key gene
(coding for the surface protein hemagglutinin) has already been completely pieced
together.
Dr. Taubenberger was kind enough to review the manuscript for "Ninth Day of
Creation" (a novel in which the 1918 strain is resurrected) and offer useful
suggestions for its improvement. The following interview with him about his group's
work was carried out through e-mail in May of 1998.
* *
* *
* * *
Interview:
CRANE: "You've indicated to me that
Ninth Day of Creation did a reasonably good
job of holding your attention.
If I can start with a question not directly related to your work, what did
you enjoy most about the book?"
TAUBENBERGER: "I liked the character development; I think the main characters
were very well developed and 'real' appearing. I also liked the suspenseful aspect
of the plot."
CRANE: "One of the plot elements in the book centers around the fictional
recovery by a biotech company of a live sample of the 1918 Spanish Flu virus.
Supposedly, in the story, the virus is isolated from a cadaver frozen for the
last eighty years on a glacier in Alaska. In real life, a Canadian-led team has
proposed visiting a cemetery located amid the permafrost of Longyearbyen, Norway,
with a similar goal in mind. You have said elsewhere that, because flu-virus is
so delicate, the chances are rather slim that the 1918 strain could exist in a
type of frozen slumber for that length of time. As a cellular pathologist, can
you foresee any conditions under which an intact sample of the Spanish flu virus
might survive to the present day?"
TAUBENBERGER: "I see no chance for the virus to survive under any natural
conditions. Permafrost fluctuates between about -10 C and +4 C, which is a
temperature zone ideally suited to kill influenza viruses. [To have survived
the last eighty years intact] would have required a sample to be frozen
continuously in liquid nitrogen, or be stored in a -70 C freezer. I do not think
either things existed in 1918. So in real life, I see no way to get live virus.
However, I have no problems with the recovery of live virus in the context of a
work of fiction---just like I had no problems with the fictional making of
dinosaurs from DNA bits in amber."

Not a mosquito, but a lizard
preserved in Dominican amber
CRANE: "That reminds me about where I got the idea for basing the virus's
recovery on a frozen tissue sample. A piece in Scientific American�, entitled
"Jurassic Virus?", mentioned that--at least up until 1993--people had
tried in vain to recover genetic snippets of the 1918 strain from human remains.
And by 1996 your group had managed just that, recovering the first fragments of
the 1918 genome--"RNA bits"--from a lung-tissue sample stored for the
last eighty years with the Institute's collection of vintage autopsy specimens.
I understand that when your paper came out in March of
1997 it attracted a good deal of attention from both inside and outside the
scientific community. Can you tell me something about why this should be so?
If the level of public interest has been high, what do you think is the reason
for our fascination with the Spanish Flu?"
TAUBENBERGER: "There has been (and continues to be) intense media interest
in our work on the 1918 influenza virus. I guess there is something about the
story which really resonates well with the press and the public's interest. One
element is the fascination [arising from the possibility] that genetic analyses
could be performed on eighty year-old autopsy tissues. The second aspect is that
here is one of the greatest medical mysteries of all time: a lethal virus
spreading around the world killing tens of millions of people in a few months.
The third facet of interest lies in the possibility that a new influenza pandemic,
and specifically one of 1918 magnitude, could happen again.
I have often been asked what is the likelihood of a
pandemic like 1918 coming again. The answer is still unknown. What is known
is that influenza pandemics occur with surprising regularity, every 20-30 years.
The chance of a new pandemic emerging is 100%. The last one was in 1968, so the
chance of a new pandemic occuring in the near future is pretty good.
We still do not know yet why 1918 was so lethal, but
whatever the circumstances were that came together to make the strain such a
killer, it could certainly happen again. The race is to find those features
[of the virus] which will help us to predict when a newly emerging strain might
share [a similar pathogenicity]. The scare over the H5N1 strain in Hong Kong
stresses the urgency of this concern."
CRANE: "What was the breakthrough in the lab that allowed you to succeed in
recovering 'ancient RNA' ? Could the technique be stretched to older material? For
instance, the Egyptians did a wonderful job of preserving bodily remains. Any chance
of seeing the remnants of RNA-virus infections there? Or is that an incredibly
remote possibility?"
TAUBENBERGER: "Our lab spent a lot of time working out how to isolate RNA
from 'suboptimal tissues' both formalin-fixed pathology material and poorly
preserved necropsy tissue from dolphins in the 1987 and 1993 mass mortality
events on the U.S. Atlantic and Gulf of Mexico coasts, respectively.
Our interest in RNA isolation from fixed biopsy material
was to develop useful clinical molecular pathology tests using tissues for which
fresh samples were unavailable. We can now do this successfully and make diagnoses
of lymphoma and leukemia, other tumors, and many viral infections, which has a
huge impact on patient care.
In regard to the dolphin research (funded by the EPA) we
were able to sequence and characterize morbilliviruses in these tissues and help
make the link between mass dolphin mortalities and infections by viruses closely
related to canine distemper virus and human measles. [References to the
Institute's dolphin morbillivirus work can be found below]
As for the mummies, your guess that RNA extraction
would be 'an incredibly remote possibility' seems like a very good answer. DNA is
however possible and has been done by Svante Paabo and others."
CRANE: "You are due to publish a second paper shortly in which you give the
complete base sequence for the 1918 gene coding for hemagglutinin--the protein
used by influenza A virus to attach itself to the host cell. How difficult was
the work? I can imagine trying to reconstruct a dropped vase and finding the job
hardest where it came to locating the smallest pieces and recognizing where they
fit in. Do similar problems turn up in your work, or are there enough big
overlapping RNA segments to facilitate the reconstruction?"
TAUBENBERGER: "It has been a very difficult task. Ann Reid in the laboratory
has been the only person doing the benchwork on the 1918 flu project. The
hemagglutinin gene has 1778 bases and the largest fragment we can amplify is about
150 bases long. There are no larger overlapping fragments. We have to design our
PCR reactions so that the sequences overlap to cover the primer sequences. You
need to know the sequence of something to do PCR, but we don't know the sequence
for the 1918 virus. Therefore we have to design 'degenerate' primers, containing
more than one base at some positions, which allows it to amplify all H1
hemagglutinins, e.g., human, swine, bird. Therefore, for every PCR reaction, we may
only get 35-100 bases of useful sequence after 1-2 weeks work.
We were also slowed down by getting two new cases. Before
proceeding any further, we decided to confirm that the gene sequence from the first
case matched those of the two new cases. It is basically the same in all three,
with only very minor changes."
CRANE: "Now that the coding for the hemagglutinin gene is known, it should be
possible to reengineer a strain of influenza virus which expresses the 1918 surface
protein. By adding genes one by one, it is also possible to imagine recreating the
original strain by the time the sequencing effort is completed. Can you foresee the
day when that might literally be accomplished?
Science derives a large measure of its power through the
sharing of data and open debate. But in a case like this--where a killer virus is
involved--it would be hard to imagine you didn't have some reservations about
publishing all your work. What are your thoughts on this issue, speaking either as
a scientist or from a personal point of view (if they aren't in complete accord
with each other)?"
TAUBENBERGER: "It is certainly possible to make recombinant influenza viruses
containing partial or complete 1918 sequences. While it does concern us somewhat,
we feel that the positive aspects of the work--finding out where the 1918 strain
came from, and why it was so lethal--far outweigh any concerns. Now that the
complete hemagglutinin sequence is known, a protective vaccine can (and will) be
made. Making such recombinant viruses may be necessary in controlled lab settings
to understand the behaviour of the 1918 virus."
CRANE: "Where do you see your work going?"
TAUBENBERGER: "We are going to generate the complete gene sequences of several
more 1918 flu genes, including neuraminidase, nucleoprotein, matrix and
nonstructural genes. This will help us get a handle on the two big issues: Where did
it come from? and, Why was it lethal?
Did it come directly from birds to humans as in Hong Kong
in 1997, or did it go through an intermediary like swine? The hemagglutinin
sequence already begins to shed light on these questions, but each gene needs to be
examined to see the complete picture emerge. Unfortunately, this will take a long
time, several years at least."
End of interview
Leonard Crane would like to express his
appreciation to Dr. Jeffery Taubenberger for taking the time to participate in this
interview.
Follow up:
When this interview was first published in 1998, work had already begun
on the reconstruction of the 1918 Spanish Flu virus by Dr. Jeffery Taubenberger
and his team at the Armed Forces Institute of Pathology. In October of 2005 it was reported
that the virus had been finally reconstructed after 10 years of work. When I interviewed him
I had forseen the day when this would happen. You can read about the announcement
in the PBS article entitled 1918 Spanish Flu Offers Clues About Pandemic Viruses.
References:
� "Initial Genetic Characterization of the 1918 'Spanish' Influzenza Virus,"
Jeffery K. Taubenberger, Ann H. Reid, Amy E. Krafft, Karen E. Bijwaard and Thomas G.
Fanning. Science, Vol. 275, 1793--1796, 1997.
� "Jurassic Virus?" Philip E. Ross. Scientific American, October 1993.
"The Flu Hunters," Erik Larson. An account of the Hong Kong Incident. Time,
February 23, 1998.
"The Dead Zone," Malcolm Gladwell. An account of modern attempts to recover the
virus responsible for the Spanish flu of 1918. The New Yorker, September 29, 1997.
"Postmortem diagnosis of Morbillivirus infection in bottlenose dolphins (Tursiops
truncatus) in the Atlantic and Gulf of Mexico Epizootics by a polymerase chain
reaction-based assay," A. Krafft, J.H. Lichy, T.P. Lipscomb, B.A. Klaunberg, S.
Kennedy and J.K. Taubenberger. J. Wildlife Diseases, Vol. 31, 410--415, 1995.
"Morbilliviral epizootic in Atlantic bottlenose dolphins of the Gulf of Mexico,"
T.P. Lipscomb, S. Kennedy, D. Moffett, A. Krafft, B.A. Klaunberg, J.H. Lichy, G.T. Regan,
G.A.J. Worthy and J.K. Taubenberger. J. Vet. Diagn. Invest. Vol. 8, 283--290, 1996.
"Two different morbilliviruses implicated in bottlenose dolphin epizootics,"
J.K. Taubenberger, M.M. Tsai, A.E. Krafft, J.H. Lichy, A.H. Reid, F.Y. Schulman and T.P.
Lipscomb. Emerging Infec. Dis. Vol. 2, 213--216, 1996
"Reevaluation of the 1987-88 Atlantic coast bottlenose dolphin (Tursiops truncatus)
mortality event with histolgic, immunohistochemical, and molecular evidence for a
morbilliviral etiology," F.Y. Schulman, T.P. Lipscomb, D. Moffett, A.E. Krafft, J.H.
Lichy, M.M. Tsai, J.K. Taubenberger and S. Kennedy. Veter. Pathol. Vol. 34,
288--295, 1997
|
Content and Copyright � by
Leonard Crane, 1998-2006.
All rights reserved.URL: http://www.ninthday.com/tauben.htm |
|
|
|


